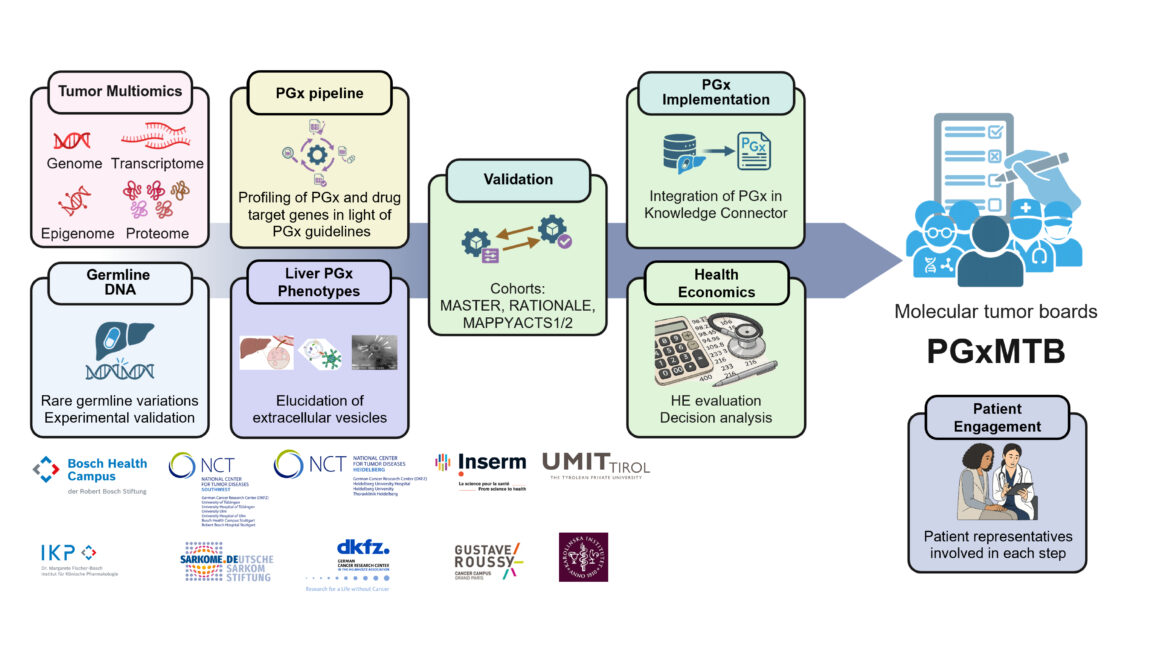
Personalised oncology has led to the development of drugs tailored to specific genetic variants. However, the full potential of targeted treatments remains untapped, largely because current approaches focus predominantly on acquired, tumour-specific alterations. Yet adverse drug reactions affecting organ systems are typically influenced by the patient’s germline rather than the tumour genome. To realise the full potential of personalised therapy, cancer care must therefore integrate pharmacogenomic (PGx) principles alongside tumour-informed strategies. PGxMTB accomplishes this mission by implementing PGx into molecular tumour boards (MTBs). Specifically, existing data from two nationwide precision oncology initiatives (DKFZ/NCT/DKTK MASTER, MAPPYACTS1/2) will be leveraged. Such implementations rely on validated computational pipelines that integrate germline PGx analysis with tumour-derived data. Building on this, a prototypic bioinformatics PGx pipeline will be refined by integrating over 300 pharmacogenes relevant for cancer and supportive care medicines. Health economic assessments will evaluate PGx implementation across different healthcare systems. In addition, the consortium aims to comprehensively map the PGx landscape in cancers. Tumour entity-specific PGx expression patterns and multiomics profiles will be analysed to uncover novel somatic PGx information, complemented by drug response data. Tumour-related changes in individual liver PGx phenotypes will be monitored using innovative liquid biopsy approaches that analyse liver-derived extracellular vesicles from cancer patients. PGxMTB brings together experts in basic and clinical PGx, precision oncology, bioinformatics, computational science, health economics, and patient engagement. Patient representatives will participate as research partners at all levels. Collectively, PGxMTB is expected to advance PGx-driven precision cancer medicine and improve patient outcomes.

