The ERKetype project focuses on cancers where a key growth and survival pathway in the cell, the RAS/RAF/MEK/ERK (ERK) signalling pathway, is abnormally active. In many patients, treatments chosen on the basis of a single gene mutation do not work well or stop working, because the cancer can bypass the blocked target or switch on alternative routes.
ERKetype aims to create “digital twins” of individual patients’ tumours. These are computer models that combine information from tumour DNA and RNA, detailed protein measurements and drug tests performed on patient-derived tumour organoids – tiny 3D cultures grown from each patient’s biopsy. By integrating these multi-omics data with mechanistic models of ERK and related signalling pathways, the digital twins are designed to predict which targeted drugs or combinations are most likely to benefit each patient.
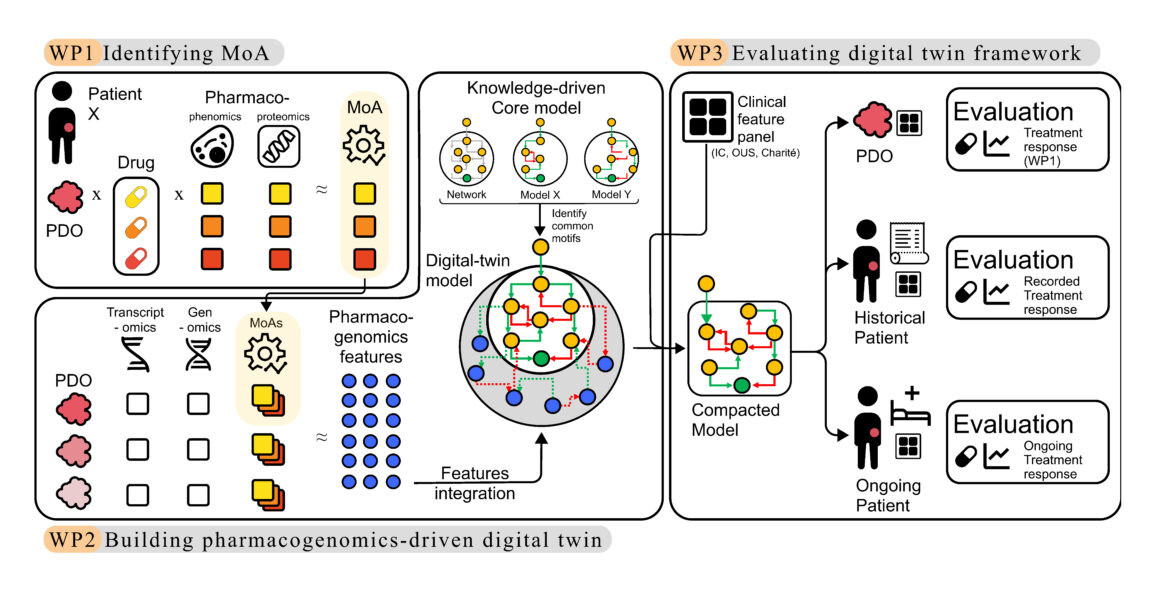
The project is structured in three main steps: mapping how drugs affect organoids at cellular and protein level; building and refining the digital twin models using pharmacogenomics data; and validating the approach using data from molecular tumour boards and precision medicine trials such as IMPRESS-Norway, SHIVA and Charité studies. The long-term goal is to provide a cost-effective decision-support tool that improves treatment selection and outcomes for patients with ERK-driven cancers.

