Stroke is the world’s second leading cause of death and the foremost cause among women. It is also a major source of long-term disability. Current acute treatments – intravenous thrombolysis with rtPA or TNK, and mechanical thrombectomy – are highly effective, yet a significant minority of patients (around 5–10%) experience severe haemorrhagic transformations (HT). These secondary brain haemorrhages, classified as PH-1 or PH-2, are associated with markedly increased mortality and disability. This project aims to develop a multi-omics prediction score capable of identifying patients at high risk of HT, thereby enabling clinicians to adjust treatment strategies and reduce the likelihood of drug-related complications.
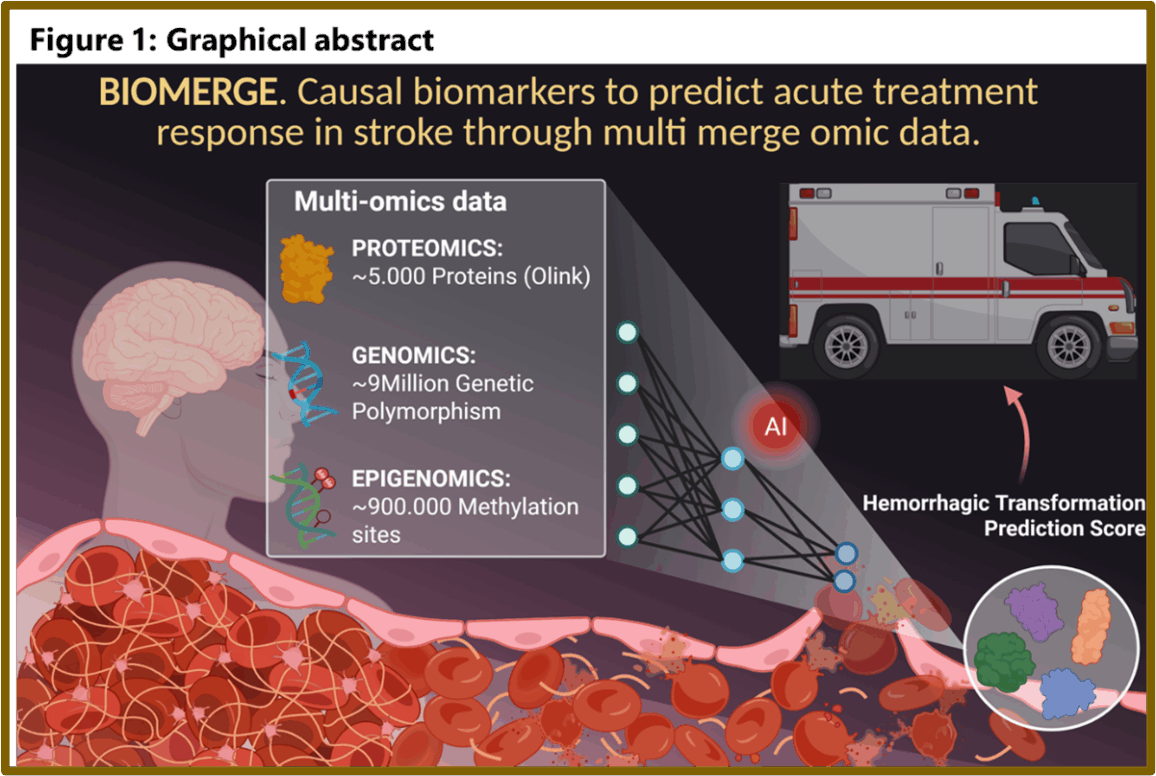
To achieve this, the study will analyse a well-defined cohort of 450 patients treated with rtPA or TNK, comparing 225 individuals who developed HT with 225 matched controls. Researchers will perform large-scale proteomic profiling of approximately 5,000 proteins, assess around nine million genetic variants, and measure methylation levels across 800,000 CpG sites. Integrative bioinformatic platforms such as Panelomix and Mergeomics will be used to combine genomic, epigenomic, and proteomic layers, helping to identify protein biomarkers suitable for rapid, point-of-care detection in ambulances.
Causal relationships between clinical factors and omics signatures will be examined using Mendelian Randomisation, supported by external datasets including the UK Biobank, ROS/MAP, and BIOMERGE. Machine-learning models will then be developed to generate a robust, generalisable prediction score that accounts for sex, gender, and ethnic diversity. The score will be prospectively validated in an independent cohort of 450 stroke patients and complemented by pharmacoeconomic analyses to assess its cost-effectiveness for real-world clinical implementation.

