EP
PerMed funded Projects
|
|
BIOKID-PGx 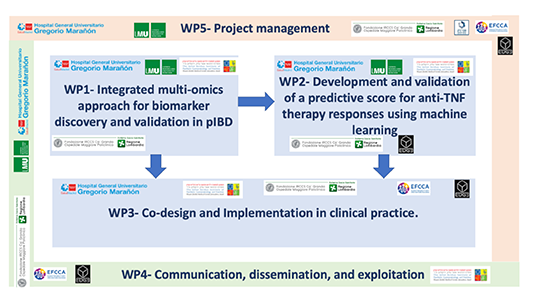
Implementation
of a multi-omic score to predict response to biological
therapy in pediatric inflammatory bowel disease
BIOKID-PGx
aims to revolutionise paediatric inflammatory bowel
disease (pIBD) treatment by pioneering personalised
medicine through an innovative, multi-omics-based risk
score predicting anti-TNF therapy failure. This
ambitious project utilises multi-omics data and
cutting-edge machine learning to identify and validate
key biomarkers associated with anti-TNF drug
response.
Read more
|
|
BIOMERGE 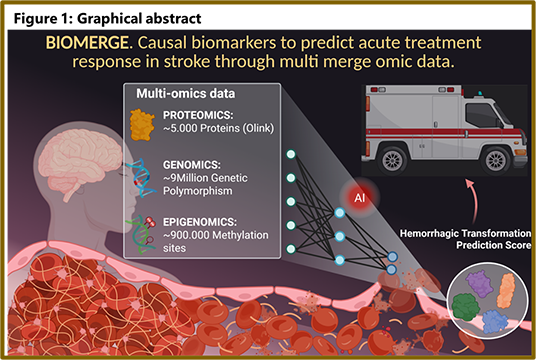
Causal
biomarkers to predict acute treatment response in stroke
through multi merge omic data.
Stroke
is one of the main causes of death and one of the main
causes of disabilities in adults. The BIOMERGE project
aims to predict the response to the treatments used
in the acute phase of stroke in order to improve the
management of the patients with a personalised medicine
and consequently decrease the disabilities and mortality
associated with this disease.
Read more
|
|
BioPATH-SAID
Pathway-Driven
Biomarker Discovery for Guiding Precision Therapy in
Undefined Systemic Autoinflammatory Diseases
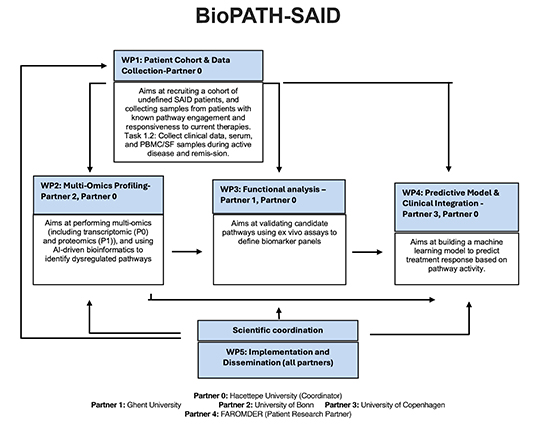
The
BioPATH-SAID project will develope an
innovative platform for pathway-driven biomarker
discovery. This platform will leverage molecular
profiling and systems biology to identify predictive
biomarkers of therapeutic response. By integrating these
biomarkers into clinical decision-making, we aim to
enable precision therapy for patients with undefined
SAIDs, improving outcomes and reducing disease
burden.
Read more
|
|
BlaCaPOx 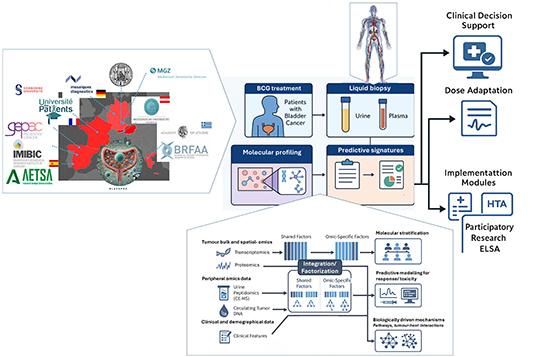
Pharmaco-omics
strategies to guide personalised medicine in BCG-
treated patients with Bladder Cancer
The
BlaCaPOx project aims to develop a novel
“pharmaco-omics” approach that uses multiple types of
biological information – such as genes, proteins, and
immune markers – to guide personalised treatment for
patients diagnosed with Bladder Cancer.
Read more
|
|
CONFUCIUS 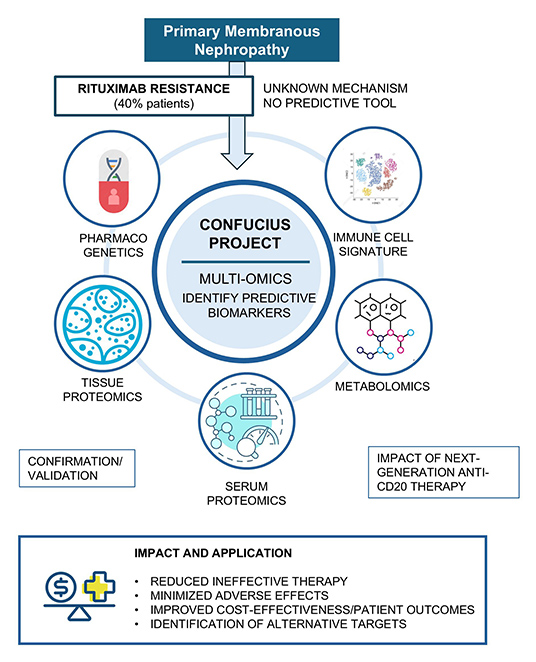
Identification
of a pharmacogenomic signature for anti-B cell precision
therapy in membranous nephropathy
The
CONFUCIUS project aims to establish a personalised
medicine framework for membranous nephropathy (MN)
through a multi-omics approach. Genetic variants, serum
and kidney proteomic profiles, and serum metabolomic
profiles will be analysed in a well-characterized
retrospective patient cohort to identify biomarkers
capable of predicting whether a patient will respond to
rituximab, thereby enabling more accurate treatment
selection. Immune cells will also be examined at
single-cell resolution to determine cellular features
associated with resistance.
Read more
|
|
EPIPREDIA 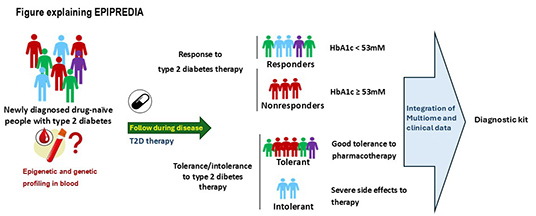
A
pharmacoepigenetic biomarker tool for prediction of
response and tolerance to treatment in type 2
diabetes
No
biomarkers have been approved for predicting response or
tolerance to Type 2 diabetes therapies. EPIPREDIA will
tackle this unmet need using our unique expertise in
epigenetics, clinical medicine and epidemiology together
with exclusive cohorts.
Read more
|
|
ERKetype 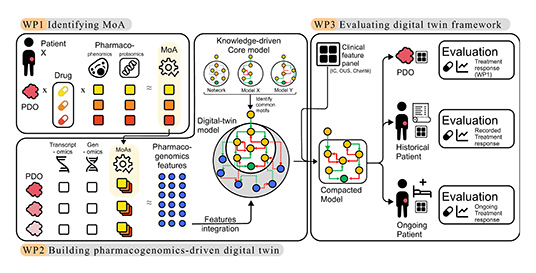
Developing
digital twins as complex biomarkers for ERK signalling
therapy through the integration of pharmaco-genomics
data and comprehensive analysis of ERK and alternative
pathway signalling dynamics
The
ERKetype project focuses on cancers where a key growth
and survival pathway in the cell, the RAS/RAF/MEK/ERK
(ERK) signalling pathway, is abnormally active. ERKetype
aims to create “digital twins” of individual patients’
tumours.
Read more
|
|
HOPE 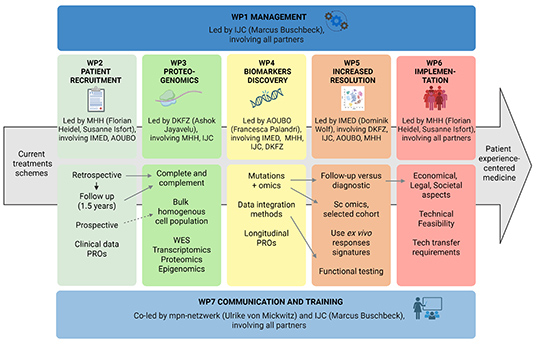
Leveraging
multidimensional high-content data for improved
patients’ experience-centred treat-ment selection for
MPN
HOPE aims
to help patients with myeloproliferative neoplasms (MPN)
receive more personalised and effective treatment. This
project will use advanced genetic and molecular data to
discover new biomarkers, which are tools that can help
guide decisions about which treatments are likely to
work best for each person. By combining detailed
genetic, protein, and molecular data (“omics” data) with
clinical results and patient-reported experiences, the
research team will uncover patterns that link specific
biological profiles to treatment responses and side
effects.
Read more
|
|
IMPACT-CRC 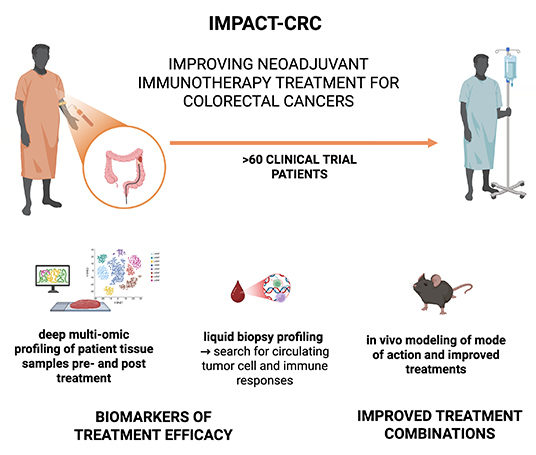
Immunotherapy
biomarkers and predictors of anti-checkpoint therapy
success in colorectal cancer
IMPACT-CRC
is tackling one of the toughest challenges in colorectal
cancer: helping patients with microsatellite-stable
(MSS) tumours benefit from immunotherapy, a treatment
that has so far worked for only a small minority of
them.
Read more
|
|
METAPRO 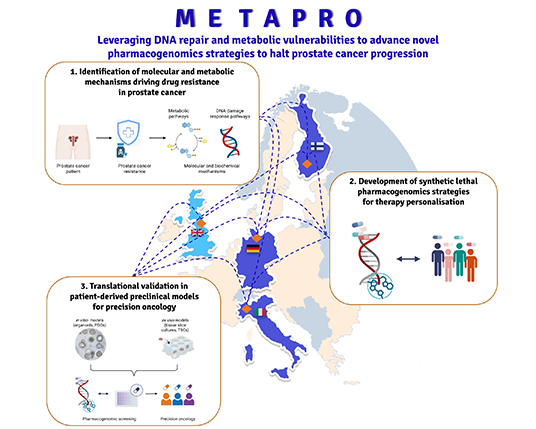
Leveraging
DNA repair and metabolic vulnerabilities to advance
novel pharmacogenomics strategies to halt prostate
cancer progression
Prostate
cancer is one of the most prevalent malignancies in men
worldwide. Although early detection and treatment of
localised disease have improved survival, metastatic
prostate cancer at diagnosis and relapses after
first-line therapies remain major clinical
challenges. The METAPRO project aims to identify
novel synthetic-lethal, pharmacogenomics-based
strategies to prevent disease progression and to improve
patient selection for targeted therapeutic
interventions.
Read more
|
|
NALUNG 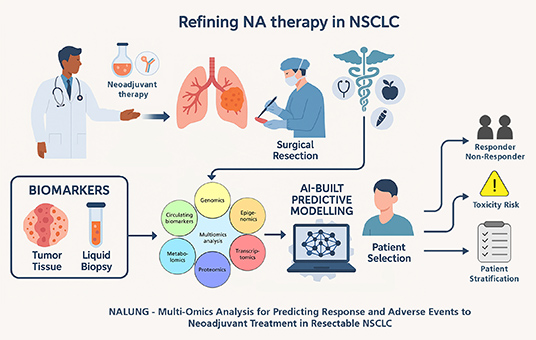
Multi-Omics
Analysis for Predicting Response and Adverse Events to
Neoadjuvant Treatment in Re-sectable NSCLC
The
NALUNG project aims to identify biological markers
in Non-Small-Cell Lung Cancer (NSCLC) tumour tissue and
blood that can reliably predict treatment response and
the risk of toxicity by applying a comprehensive
multi-omics strategy that includes genomics,
epigenomics, transcriptomics, proteomics, and
metabolomics.
Read more
|
|
NordicPCPM
Forum 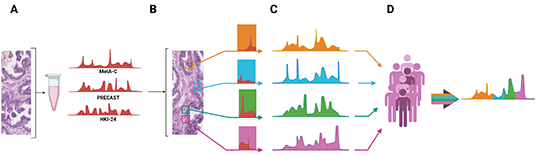
Spatially
curated multi-omic biomarkers to delineate a roadmap for
personalised medicine in prostate cancer: A study within
the Nordic Prostate Cancer Personalized Medicine Forum
(Nordic PCPM Forum)
The
project aims to develop spatially informed multiomics
biomarker panels that can more accurately predict
treatment response and disease progression in prostate
cancer. The consortium will refine and validate
biomarker signatures in well-characterised patient
cohorts and evaluate their real-world usability through
the SPRINTR clinical infrastructure. This approach will
support earlier, more personalised treatment decisions
and improve patient outcomes.
Read more
|
|
omiCSFit 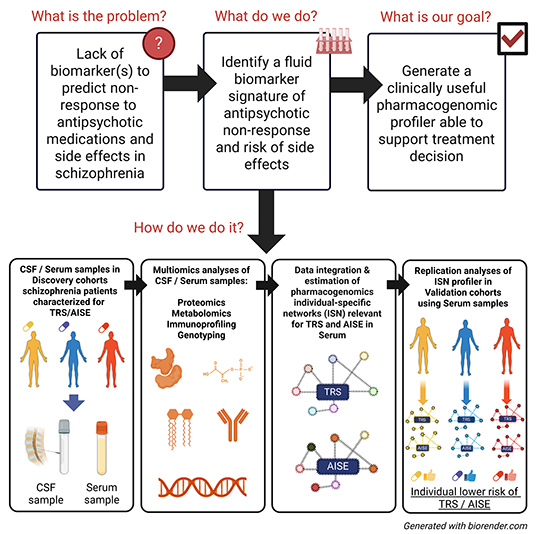
Multiomics
CSF-serum analysis in schizophrenia: characterisation of
drug-related and individual-specific networks for
personalised antipsychotic treatment
The
omiCSFit project aims to lay the groundwork for
developing support tools for clinicians for the early
prediction of TRS- and AISE-related outcomes in SCZ
patients. By utilising unique cerebrospinal fluid (CSF)
and serum samples of SCZ patients, combined with
state-of-the-art multiomics approaches and advanced
statistical techniques, the project will define
individualised drug-specific pharmacogenomic profiles
for the early prediction of TRS/AISE risk.
Read more
|
|
Optimize 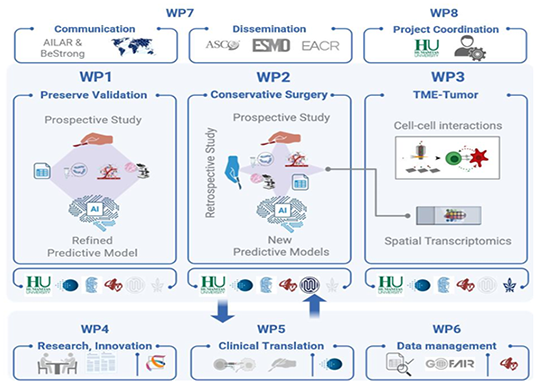
Multiomics
signature to customise induction or neoadjuvant
treatments for optimising laryngeal preservation
strategies
The
Optimize project aims to define a tailored approach to
locally advanced laryngeal/hypopharyngeal squamous cell
carcinoma (LHSCC), applying multiomic (clinical,
genomic, pathomic) signatures to refine neoadjuvant
strategies for surgical laryngeal preservation and
curative radiotherapy.
Read more
|
|
PGxMTB 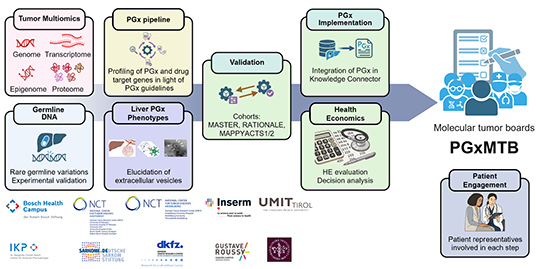
Implementation
of Pharmacogenomics (PGx) into Molecular Tumour Boards
(PGxMTB) in Europe: Improved prediction of PGx
phenotypes using germline and tumour multiomics
data
PGxMTB
addresses an unmet need in personalised cancer care by
integrating pharmacogenomics (PGx) knowledge and tools
beyond tumour DNA-level information into the clinical
setting of molecular tumour boards (MTB) to minimise
adverse drug reactions and avoid ineffective cancer and
supportive treatments.
Read more
|
|
PHARAO 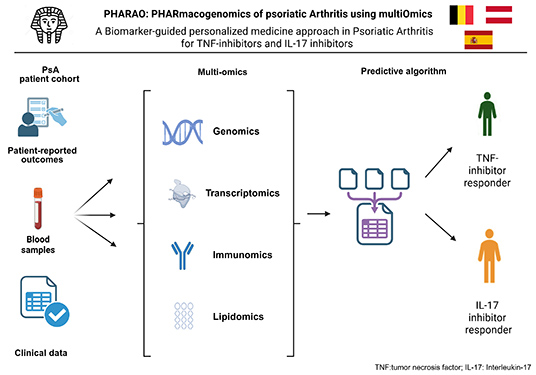
PHARmacogenomics
of psoriatic Arthritis using multiOmics
Psoriatic
arthritis (PsA) is a chronic inflammatory disease
affecting the joints and skin, often leading to
progressive joint damage and impaired quality of life.
The PHARAO consortium aims to develop a predictive tool
that identifies which PsA patients will respond best to
specific biological treatments, enabling personalised
therapy.
Read more
|
|
PhORECaST 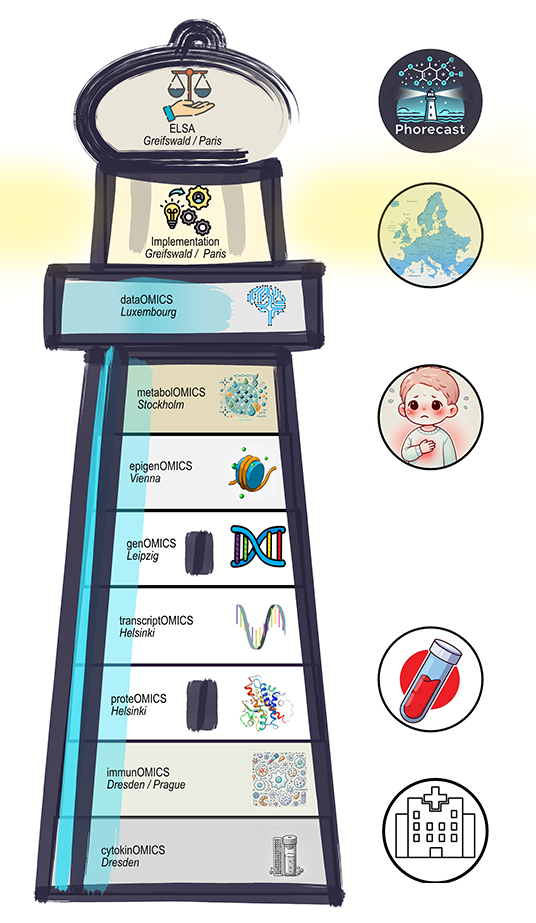
PharmocogenOmics
for minimised Risk and better Efficacy in Children on
highdose Steroid Treatment
The
PhORECaST project will conductia multi-centre,
multi-omics study to compare the clinical efficacy and
long-term outcomes of a 3-day vs. 5-day pulse of
high-dose glucocorticoids in children with acute
immunemediated disorders. The goal is to identify
biomarkers that predict individual response and risk of
toxicity, and to lay the groundwork for evidence-based
treatment algorithms integrating alternative targeted
therapies when glucocorticoids are likely to be
ineffective or harmful.
Read more
|
|
pRCC-TREAT 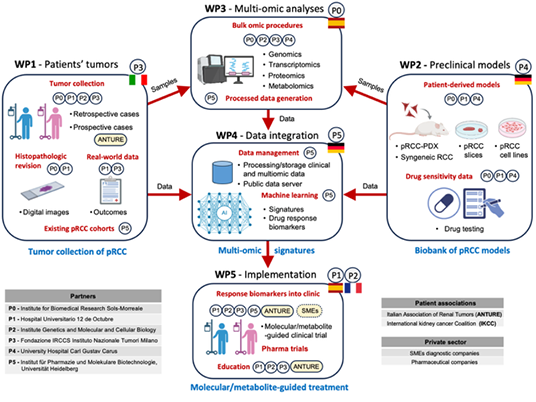
Papillary
renal cancer, metastatic patients, molecular
stratification, multiomics, data integration, immune
checkpoint inhibitors, antiangiogenic drugs
The
goal of the pRCC-TREAT project is to understand the
molecular and metabolic makeup of metastatic papillary
renal cell carcinoma (pRCC), identify biomarkers that
predict which treatments will work, and discover new
drug targets. The project will also create the world’s
largest research resource for metastatic pRCC, including
tumour samples, laboratory models, and a detailed
clinical and molecular database.
Read more
|
|
PRECISION-tMN 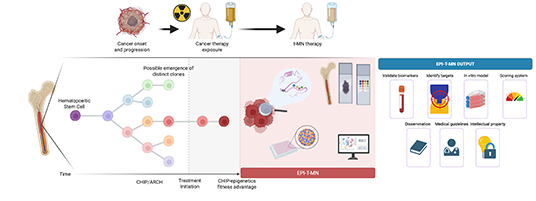
Precision
Medicine for Therapy-Related Myeloid Neoplasias
PRECISION-tMN
aims to revolutionise the clinical management of
therapy-related myeloid neoplasms (t-MN) by integrating
precision medicine approaches with cutting-edge
molecular and computational analyses.
Read more
|
|
SPOCK 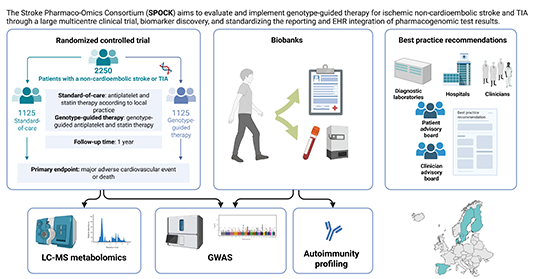
Stroke
Pharmaco-Omics Consortium
Stroke
remains a leading cause of death and a significant
contributor to the global disease burden, with over
840,000 affected annually in the EU. The SPOCK
project aims to improve the management of ischemic
non-cardioembolic stroke and transient ischemic attack
(TIA) using genotype-guided therapy.
Read more
|
|
STAR-MBM 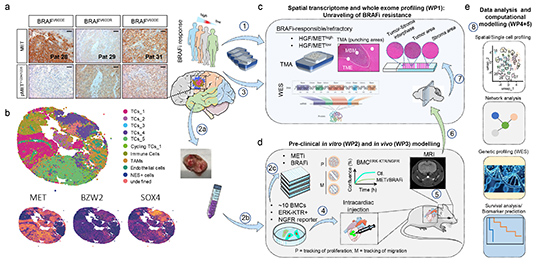
Strategy
to Target Acquired Resistance in Melanoma Brain
Metastases
The
STAR-MBM project’s main aim and the consoritum’s vision
is to combat therapy-resistance in patients with
melanoma brain metastases through spatial transcriptomic
and genetic profiling and pre-clinical and in-silico
models.
Read more
|
|
TRANS-AML-EU 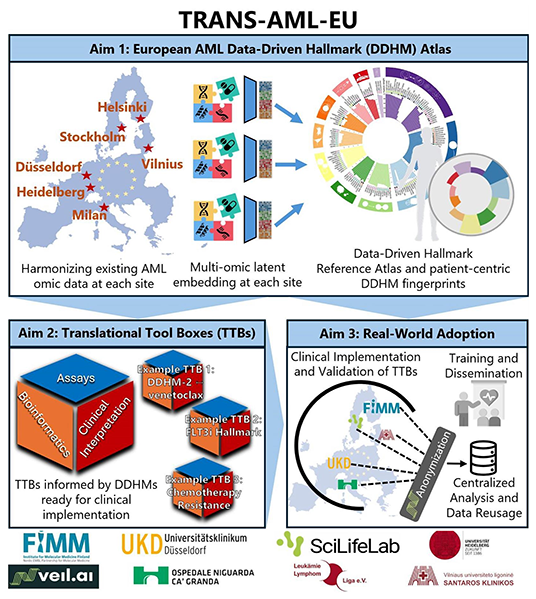
Transforming
Personalised Acute Myeloid Leukaemia Therapy through
an
Integrative European Multi-Centre Multi-Omics
Network
TRANS-AML-EU
aims to transform how therapies are selected for
patients with acute myeloid leukemia (AML). Rather than
relying on single mutations to categorise patients into
disease subgroups, the project combines genomics,
epigenomics, transcriptomics, proteomics and ex vivo
drug-response profiling to derive latent disease
programs, termed Data-Driven Hallmarks (DDHMs).
Read more
|
|
See all projects
|























